Note
Go to the end to download the full example code.
Grader#
This example shows how to use the Grader class to grade
configurations with the trained potential and the training configurations.
import logging
import ase.io
import matplotlib.pyplot as plt
import numpy as np
from motep.grade import Grader
from motep.io.mlip.cfg import write_cfg
from motep.io.mlip.mtp import read_mtp
If you need a log, you can set the following.
logging.basicConfig(level=logging.INFO, format="%(message)s")
We first load the trained potential and the corresponding training dataset.
images_training = ase.io.read("../0.0_train/ase.xyz", index=":")
mtp_data = read_mtp("final.mtp")
mtp_data.species = [6, 1]
We then load/make the configurations to grade. In this example, we make configurations slightly perturbed from those in the training dataset.
rng = np.random.default_rng(42)
images_in = []
for atoms_ref in images_training:
atoms = atoms_ref.copy()
atoms.rattle(0.5, rng=rng)
images_in.append(atoms)
We then make the Grader class and grade the images by the potential.
grader = Grader(mtp_data, seed=42)
grader.update(images_training)
images_out = grader.grade(images_in)
After the grading, the grades are stored in atoms.calc.results.
images_out[0].calc.results["MV_grade"]
np.float64(0.9054087181609364)
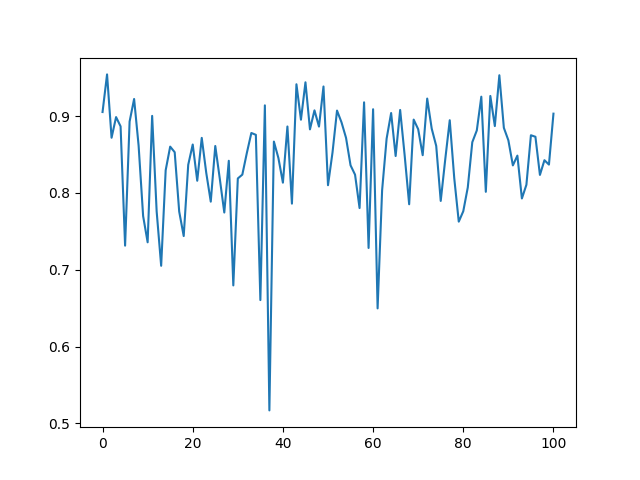
We plot the grades of the configurations.
grades = [atoms.calc.results["MV_grade"] for atoms in images_out]
plt.plot(grades)
plt.show()
We finally store the graded configurations.
write_cfg("graded.cfg", images_out)
In order to write the results in the extended-xyz format,
at this moment we need to restore the results in atoms.info
(or atoms.arrays for neighborhood grades).
for atoms in images_out:
for key in ["MV_grade"]:
atoms.info[key] = atoms.calc.results.pop(key)
ase.io.write("graded.xyz", images_out)
Total running time of the script: (0 minutes 0.352 seconds)